 DRKG is a comprehensive biological knowledge graph that relates human genes, chemical compounds, biological processes, drug side effects, diseases, and symptoms. It curates and normalizes data from six publicly available databases as well as information from recent publications related to COVID-19.
DRKG is a comprehensive biological knowledge graph that relates human genes, chemical compounds, biological processes, drug side effects, diseases, and symptoms. It curates and normalizes data from six publicly available databases as well as information from recent publications related to COVID-19.
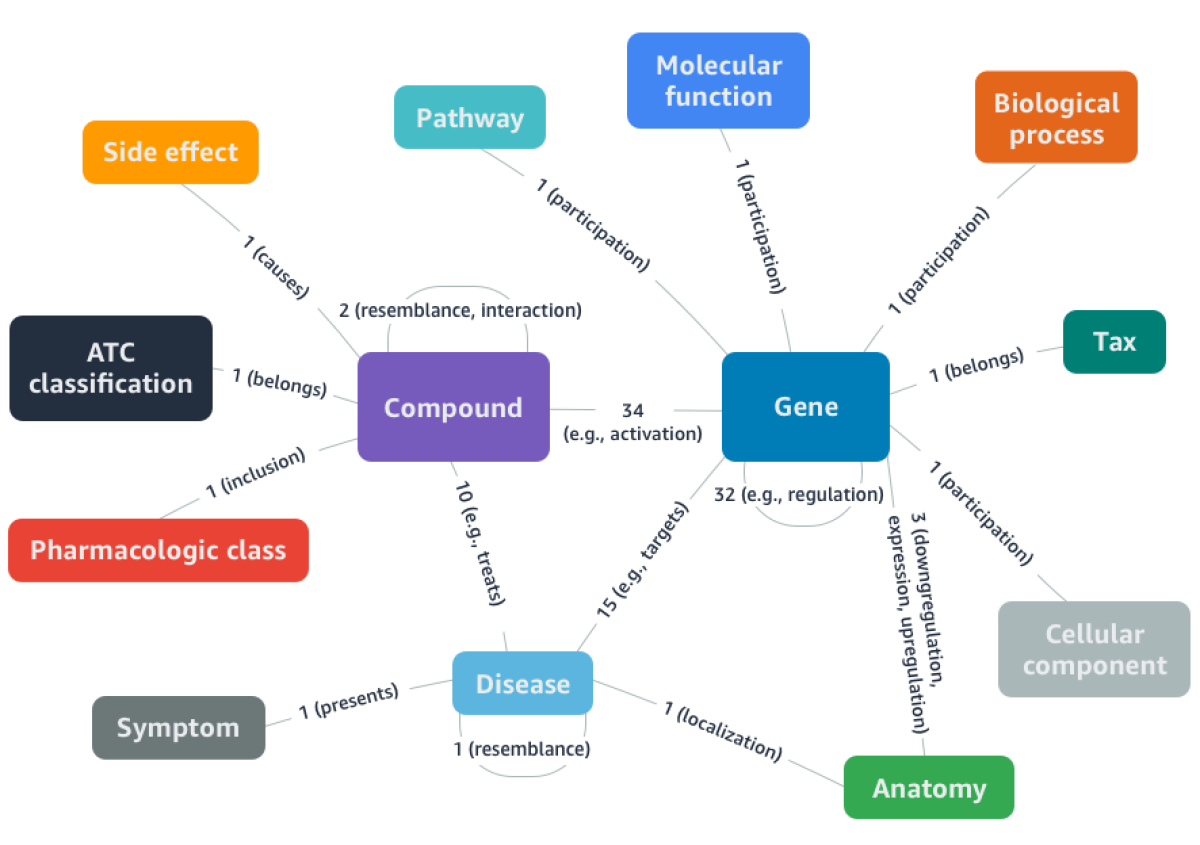
DRKG includes nearly 100,000 entities of more than a dozen types and nearly 6,000,000 relationships of more than 100 types. It captures interactions between entities that are related to the genetic signature of COVID-19 or to components of existing drugs and viruses.
The associated machine learning tools use state-of-the-art deep-graph-learning methods (DGL-KE) that take advantage of distributed graph operations (from popular deep-learning libraries such as PyTorch and MXNet) to predict the likelihood that a drug can treat a disease or bind to a protein associated with the disease.
When tested against the human proteins associated with COVID-19, these tools assigned high probabilities to many of the COVID-19 drug candidates currently in clinical trials. Both DRKG and the machine learning tools are publicly available on GitHub. This should help make computational drug repurposing for COVID-19 and other diseases (e.g., Alzheimer’s disease) more efficient and effective.