At the Sullivan Lab, we have been developing new phage-host model systems with phages infecting heterotrophic bacteria (heterophages), as well as advancing more well-established model systems with phages infecting marine cyanobacteria (cyanophages).
Navigate to:
Cyanophages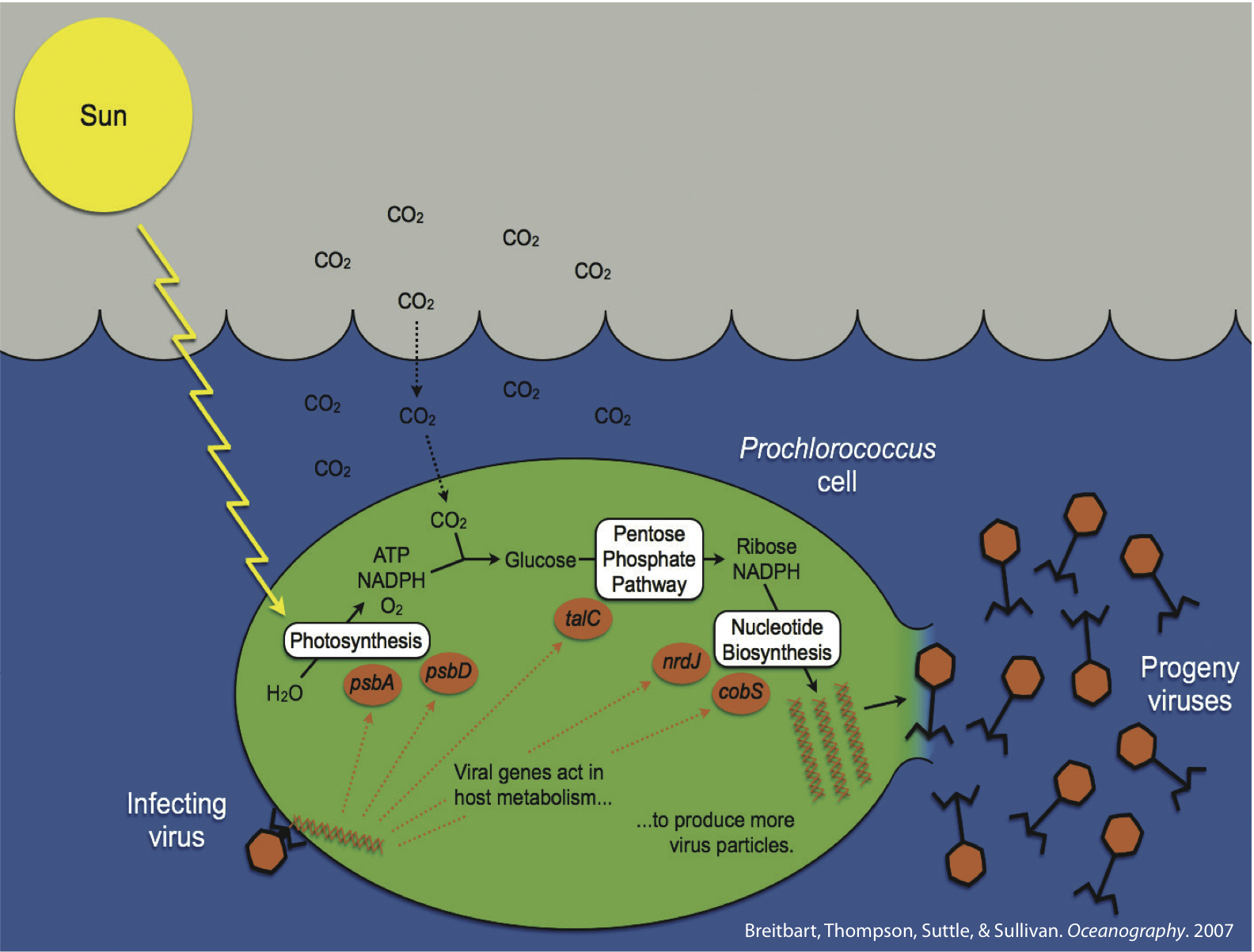
The marine cyanobacteria Prochlorococcus and Synechococcus are globally important primary producers. In spite of their small size, these cyanobacterial cells are numerically dominant over vast areas of the “desert oceans” and are significant contributors to global carbon cycling. These were some of the earliest “ecological microbes” sequenced by the DOE JGI, which led to an understanding of the genomic underpinnings that are responsible for their ecological niches in the environment
Cyanobacterial viruses (cyanophages) are abundant, contribute to host mortality, and impact marine cyanobacterial diversity through mortality and moving genes through the host population. Cyanophage genomes appear to be standard “coliphage” decorated with a suite of niche- and host-defining genes. These latter genes include photosynthesis genes, as well as genes likely involved in carbon metabolism, phosphate stress and novel nucleotide metabolism genes. Detailed studies of the core reaction center genes suggest that they are widespread among cyanophage isolates with a seemingly predictable distribution, they are expressed during infection, and parts of the viral copies of the genes have even been transferred back into their hosts – thus cyanophages are acting as evolutionary drivers of the core reaction centers of the numerically dominant photosystems on the planet.
Heterophages
The role of heterotrophic bacteria, such as Cellulophaga and Pseudoalteromonas, and their phages has been underexplored in the wild. We are interested in using genomically-characterized representative host strains to develop novel phage-host model systems that can be used for hypothesis testing in non-photosynthetic microbes.

Comments are closed, but trackbacks and pingbacks are open.